

Santa Monica College
Welcome to the AIAA SMC Student Initiative — where students unite to learn, build, and innovate in aviation and space exploration.
Who We Are
The AIAA SMC Student Initiative is a student-led organization passionate about all things aerospace. We collaborate on hands-on projects, connect with industry professionals, and inspire the next generation of engineers and explorers.
Astrophysics Research
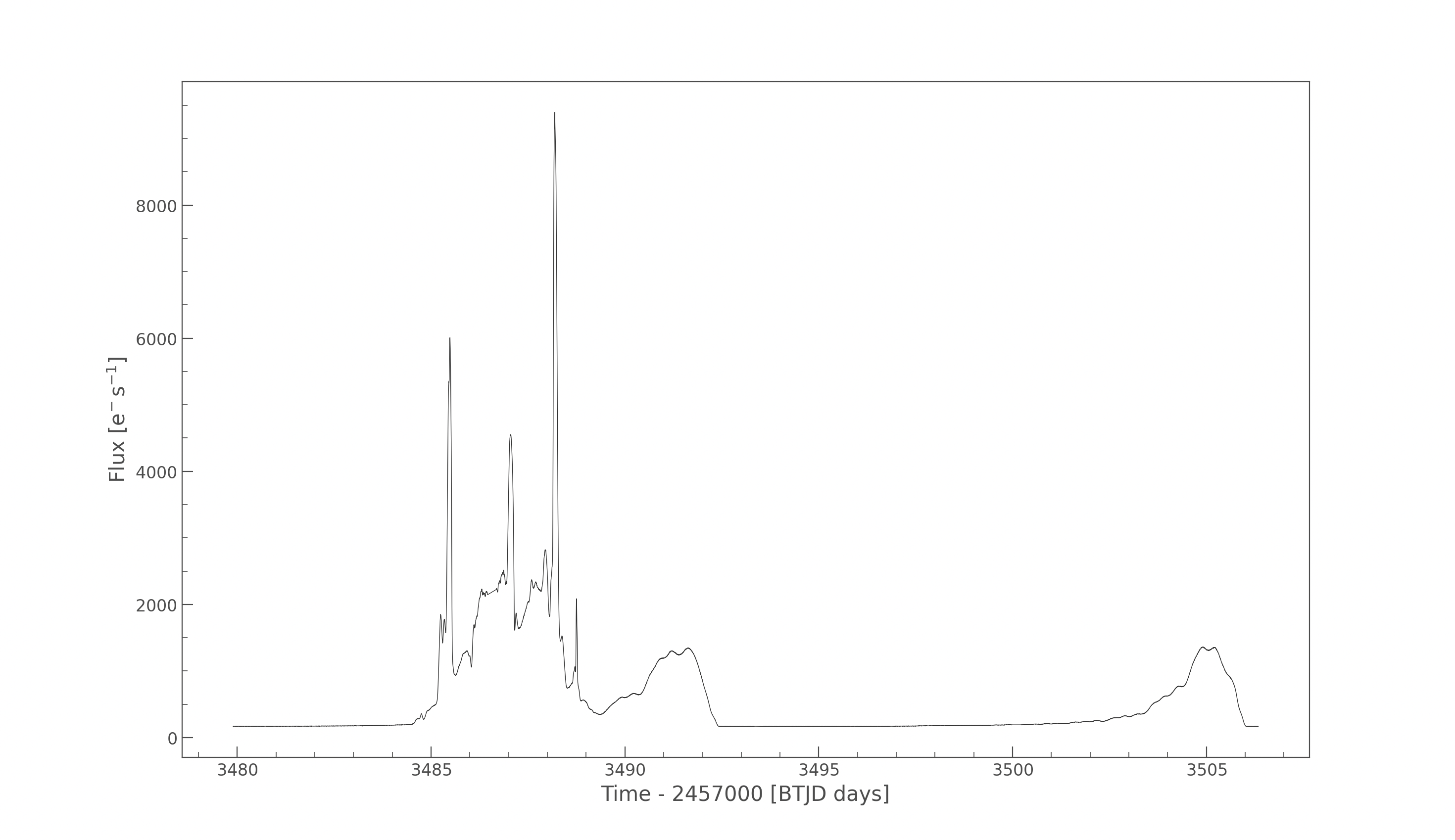
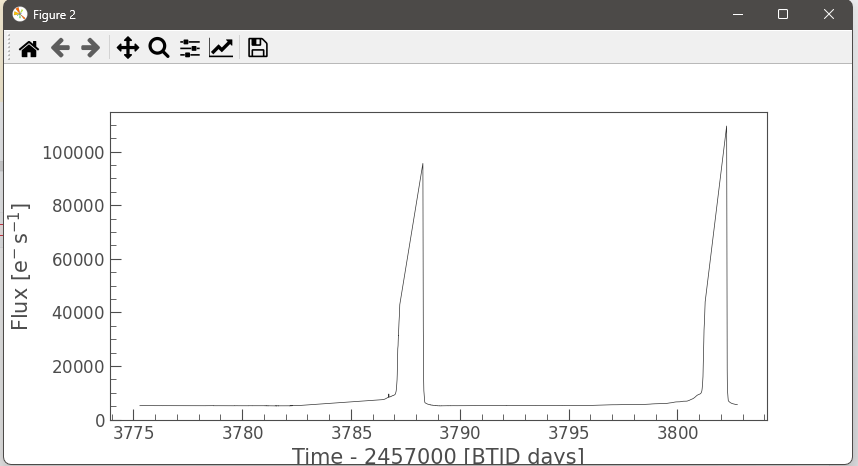
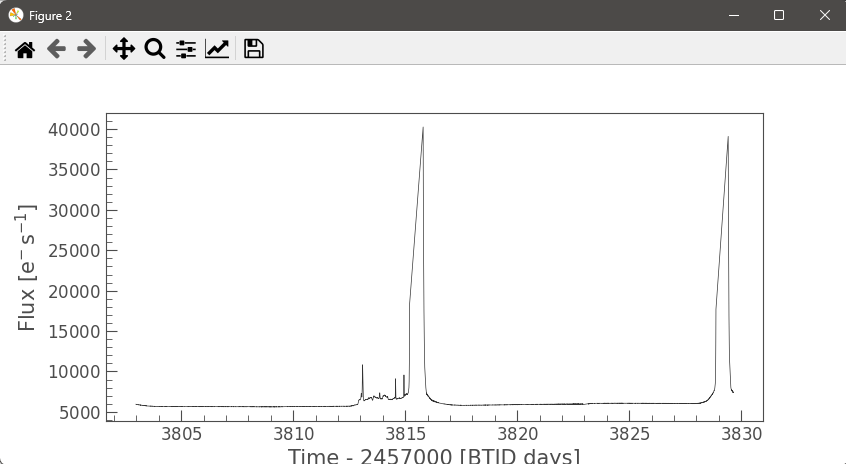
Our members are working with real TESS satellite data to detect and characterize stellar flares and study brightness variations of celestial objects over time. The pipeline builds aperture-masked light curves, computes flux-weighted centroids, and measures centroid shifts to distinguish on-target flares from nearby contaminants.
An upcoming large-scale extension will survey hundreds of stars in the NGC 2516 open cluster using Gaia DR3 membership data and automated anomaly detection.
Competition · 2025–2026
We are competing in the AIAA Undergraduate Team Space Design Competition — a national challenge where student teams develop a full mission concept and proposal. Read the brief below, and track our upcoming deliverables.
Read the full competition requirements, challenge statement, and submission guidelines from AIAA.
Team Deliverables
Active Research
Explore the actual Python code powering our astrophysics research — from single-star flare detection to large-scale open cluster surveys.
"""
flare_ch.py
-----------
Detects and characterizes stellar flares from TESS Target Pixel File (TPF) data.
Pipeline overview:
1. Load a TESS TPF cutout and generate an aperture-masked light curve.
2. Identify the brightest cadence (strongest flare candidate).
3. Compute flux-weighted centroids over time using only aperture pixels.
4. Measure the centroid shift between the pre-flare baseline and the flare peak.
5. Project the target star's sky coordinates onto the pixel grid via WCS.
A significant centroid shift during the flare suggests the emission originates
from a nearby contaminating source rather than the target star itself.
"""
from lightkurve import TessTargetPixelFile
import matplotlib.pyplot as plt
import numpy as np
from astropy.coordinates import SkyCoord
import astropy.units as u
# ---------------------------------------------------------------------------
# 1. Load data
# ---------------------------------------------------------------------------
tpf_path = r"/Users/noel/Downloads/AIAA/Star Project/tess-s0080-2-2_270.420554_28.932725_30x30_astrocut.fits"
tpf = TessTargetPixelFile(tpf_path)
# ---------------------------------------------------------------------------
# 2. Build aperture mask and light curve
#
# The same mask is reused for the light curve, flare detection, and centroid
# computation to ensure all three are internally consistent.
# Pixels with flux > 3x the background median are included.
# ---------------------------------------------------------------------------
aperture_mask = tpf.create_threshold_mask(threshold=3)
lc = tpf.to_lightcurve(aperture_mask=aperture_mask)
plt.figure(figsize=(10, 4))
lc.plot()
plt.show()
# ---------------------------------------------------------------------------
# 3. Extract raw flux and time arrays
# ---------------------------------------------------------------------------
flux = tpf.flux.value # shape: (n_cadences, ny, nx)
time = tpf.time.value # shape: (n_cadences,) — TBJD
print("Flux shape:", flux.shape)
print("Time len: ", len(time))
# ---------------------------------------------------------------------------
# 4. Apply aperture mask and detect the strongest flare
#
# Non-aperture pixels are set to NaN so they contribute nothing to sums.
# The cadence with the highest total in-aperture flux is taken as the flare peak.
# ---------------------------------------------------------------------------
masked_flux = flux.copy()
masked_flux[:, ~aperture_mask] = np.nan # blank out background pixels
aperture_flux = np.nansum(masked_flux, axis=(1, 2)) # total in-aperture flux per cadence
flare_idx = np.nanargmax(aperture_flux) # index of peak brightness
print("Index of strongest flare:", flare_idx)
print("Time of strongest flare: ", time[flare_idx])
# ---------------------------------------------------------------------------
# 5. Compute flux-weighted centroids at every cadence
#
# Centroid formula: x_c = sum(x * F) / sum(F)
# Computed only over aperture pixels; NaNs are excluded by nansum.
# ---------------------------------------------------------------------------
ny, nx = flux.shape[1], flux.shape[2]
y, x = np.mgrid[0:ny, 0:nx] # pixel coordinate grids
num_x = np.nansum(x * masked_flux, axis=(1, 2)) # sum(x * F)
num_y = np.nansum(y * masked_flux, axis=(1, 2)) # sum(y * F)
den = np.nansum(masked_flux, axis=(1, 2)) # sum(F)
centroid_x = num_x / den # flux-weighted x centroid at each cadence
centroid_y = num_y / den # flux-weighted y centroid at each cadence
# ---------------------------------------------------------------------------
# 6. Select pre-flare baseline index
#
# Prefer 10 cadences before the peak. If the flare occurs within the first
# 10 cadences, fall back to 10 cadences after the peak.
# NaN frames are skipped in either direction.
# ---------------------------------------------------------------------------
if flare_idx >= 10:
pre_idx = flare_idx - 10
while pre_idx > 0 and np.all(np.isnan(masked_flux[pre_idx])):
pre_idx -= 1
else:
pre_idx = flare_idx + 10
while pre_idx < len(time) - 1 and np.all(np.isnan(masked_flux[pre_idx])):
pre_idx += 1
print("Warning: flare near start of data; using post-flare frame as baseline")
peak_idx = flare_idx
while peak_idx > 0 and np.all(np.isnan(masked_flux[peak_idx])):
peak_idx -= 1
print("Using pre-flare index:", pre_idx, "time:", time[pre_idx])
print("Using peak index: ", peak_idx, "time:", time[peak_idx])
# ---------------------------------------------------------------------------
# 7. Measure centroid shift between baseline and flare peak
# ---------------------------------------------------------------------------
x_pre = centroid_x[pre_idx]
y_pre = centroid_y[pre_idx]
x_peak = centroid_x[peak_idx]
y_peak = centroid_y[peak_idx]
print("Centroid BEFORE flare: x =", x_pre, " y =", y_pre)
print("Centroid DURING flare: x =", x_peak, " y =", y_peak)
dx = x_peak - x_pre
dy = y_peak - y_pre
print("Centroid shift (dx, dy) =", dx, dy)
# ---------------------------------------------------------------------------
# 8. Project target star coordinates onto the pixel grid via WCS
# ---------------------------------------------------------------------------
wcs = tpf.wcs
coord = SkyCoord(ra=270.420554*u.deg, dec=28.932725*u.deg) # TIC 1603615314
x_tic, y_tic = wcs.world_to_pixel(coord)
print("TIC 1603615314 pixel position: x =", x_tic, " y =", y_tic)"""
Variable Star Anomaly Detection Pipeline
Target: NGC 2516 open cluster (~400-500 members)
Phase 1: Gaia membership query + TESS light curve download
"""
import pandas as pd
from pathlib import Path
from astroquery.gaia import Gaia
from astroquery.mast import Catalogs
import lightkurve as lk
import warnings
warnings.filterwarnings("ignore")
# -- Output directory ----------------------------------------------------------
DATA_DIR = Path("data/ngc2516")
DATA_DIR.mkdir(parents=True, exist_ok=True)
# -- 1. Query Gaia DR3 membership for NGC 2516 ---------------------------------
# Center: RA=119.52, Dec=-60.75 | Distance ~408 pc -> parallax ~2.45 mas
# Membership filter: parallax + proper motion box from Cantat-Gaudin 2018
GAIA_QUERY = """
SELECT
source_id,
ra,
dec,
parallax,
parallax_error,
pmra,
pmdec,
phot_g_mean_mag,
phot_bp_mean_mag,
phot_rp_mean_mag,
bp_rp,
radial_velocity
FROM gaiadr3.gaia_source
WHERE CONTAINS(
POINT(ra, dec),
CIRCLE(119.52, -60.75, 0.75)
) = 1
AND parallax BETWEEN 2.1 AND 2.9
AND pmra BETWEEN -5.5 AND -3.5
AND pmdec BETWEEN 11.0 AND 13.0
AND phot_g_mean_mag < 14.0
AND parallax_error < 0.2
"""
print("Querying Gaia DR3 for NGC 2516 members...")
job = Gaia.launch_job(GAIA_QUERY)
gaia_members = job.get_results().to_pandas()
print(f" Found {len(gaia_members)} candidate members\n")
gaia_members.to_csv(DATA_DIR / "gaia_members.csv", index=False)
# -- 2. Cross-match Gaia sources to TESS Input Catalog (TIC) ------------------
print("Cross-matching to TESS Input Catalog...")
tic_ids = []
tic_tmag = []
for _, row in gaia_members.iterrows():
result = Catalogs.query_region(
f"{row['ra']} {row['dec']}",
radius="5s", # 5 arcsec search radius
catalog="TIC"
)
if len(result) > 0:
result.sort("Tmag") # take brightest match within radius
tic_ids.append(int(result["ID"][0]))
tic_tmag.append(float(result["Tmag"][0]))
else:
tic_ids.append(None)
tic_tmag.append(None)
gaia_members["tic_id"] = tic_ids
gaia_members["Tmag"] = tic_tmag
matched = gaia_members.dropna(subset=["tic_id"])
matched = matched[matched["Tmag"] < 13.5].reset_index(drop=True)
matched["tic_id"] = matched["tic_id"].astype(int)
print(f" {len(matched)} stars matched to TIC with Tmag < 13.5\n")
matched.to_csv(DATA_DIR / "tic_matched.csv", index=False)
# -- 3. Download TESS light curves --------------------------------------------
# NGC 2516 covered in Sectors 1, 27, 28 (Southern CVZ-adjacent)
# Priority: 2-min cadence SPOC PDCSAP; fallback to QLP (FFI-based)
TARGET_SECTORS = [100]
PREFERRED_AUTHORS = ["SPOC", "QLP"]
print("Downloading TESS light curves...")
light_curves = {}
download_log = []
for _, row in matched.iterrows():
tic_id = row["tic_id"]
target = f"TIC {tic_id}"
try:
search = lk.search_lightcurve(
target,
mission="TESS",
sector=TARGET_SECTORS,
author=PREFERRED_AUTHORS,
exptime="short" # prefer 2-min cadence
)
# Fallback: if no 2-min, accept 10-min (QLP FFI)
if len(search) == 0:
search = lk.search_lightcurve(
target, mission="TESS",
sector=TARGET_SECTORS, author="QLP"
)
if len(search) == 0:
download_log.append({"tic_id": tic_id, "status": "no_data"})
continue
lc_collection = search.download_all(
flux_column="pdcsap_flux",
quality_bitmask="hardest"
)
if lc_collection is None or len(lc_collection) == 0:
download_log.append({"tic_id": tic_id, "status": "download_failed"})
continue
# Stitch sectors: normalize each to unit median before combining
lc_stitched = lc_collection.stitch(corrector_func=lambda lc: lc.normalize())
lc_clean = (
lc_stitched
.remove_nans()
.remove_outliers(sigma=5.0)
)
if len(lc_clean) < 200:
download_log.append({"tic_id": tic_id, "status": "too_short",
"n_points": len(lc_clean)})
continue
light_curves[tic_id] = lc_clean
download_log.append({
"tic_id": tic_id,
"status": "ok",
"n_points": len(lc_clean),
"sectors": list(lc_collection.sector) if hasattr(lc_collection, "sector")
else TARGET_SECTORS,
"Tmag": row["Tmag"],
"bp_rp": row["bp_rp"]
})
except Exception as e:
download_log.append({"tic_id": tic_id, "status": f"error: {str(e)}"})
continue
# -- 4. Summary ----------------------------------------------------------------
log_df = pd.DataFrame(download_log)
log_df.to_csv(DATA_DIR / "download_log.csv", index=False)
ok = log_df[log_df["status"] == "ok"]
print(f"Download complete:")
print(f" Successfully downloaded: {len(ok)} light curves")
print(f" No data available: {len(log_df[log_df['status'] == 'no_data'])}")
print(f" Too short / failed: {len(log_df) - len(ok) - len(log_df[log_df['status'] == 'no_data'])}")
print(f"\nLight curves ready for feature extraction: {len(light_curves)}")
print(f"Data saved to: {DATA_DIR.resolve()}")Spring 2026 · Every Sunday @ 9:30 PM
Weekly sessions focused on the AIAA Space Design Competition — from onboarding through final proposal submission. All meetings held online.
Have a question or want to get involved? We'd love to hear from you. Reach out anytime — we welcome new members and collaborators at all levels.
aiaa.smc.student.branch@gmail.com
Community
Connect With Us
Stay in touch, get updates, and join the conversation online.